Peer-reviewed work.
No noise.
The publications listed below represent research conducted by members of the Boring Science team across academic and independent research environments. Built on real data, rigorous methods, and open deposition.
bsdock: An SE(3)-Equivariant Graph Neural Network Scoring Function for Molecular Docking
Preprint · Under Submission
Accurate prediction of protein-ligand binding affinity remains a central challenge in computational drug discovery. We present bsdock, a comprehensive molecular docking platform featuring a novel SE(3)-equivariant graph neural network scoring function. Protein-ligand complexes are encoded as heterogeneous graphs with physically grounded rotational and translational invariance enforced by construction. Trained on combined ULVSH (942 compounds) and PDBbind (5,316 complexes) datasets, bsdock achieves Pearson R = 0.88 on PDBbind and R = 0.82 on ULVSH, substantially outperforming AutoDock Vina, Gnina, and MM-GBSA. Activity classification yields AUC-ROC = 0.94. The platform integrates traditional pose generation with GNN rescoring in a hybrid workflow, with specialist modules for flexible, metal-coordinated, and tethered docking.
bsvision: Physics-Constrained Diffusion Models for Temporally Consistent Medical Video Generation
Preprint · Under Submission
Generating physiologically coherent synthetic medical video remains an open challenge — standard video diffusion models optimize perceptual quality without enforcing the physics of biological motion. We present bsvision, a physics-constrained pixel-space video diffusion model for echocardiography generation. Our 3D UNet backbone with cross-frame attention at all encoder/decoder levels is conditioned on cardiac cycle embeddings to enforce temporally consistent dynamics. Through five model iterations we identify that cross-frame attention (+0.32 SSIM) and SSIM loss weight tuning (+0.32 SSIM) dominate performance, while physics-based losses provide regularization (+0.04 SSIM). The final model achieves SSIM = 0.87 between consecutive frames — a 6.2× improvement over baseline — and augmenting with 10k synthetic samples yields a 12.8% improvement in downstream ejection fraction prediction.
Progenitor-associated cells in the adult human spinal cord exhibit state-dependent proliferative competence
Under Review
Integrating anatomically targeted ex vivo assays, bulk RNA sequencing, and re-analysis of adult human single-nucleus and spatial transcriptomic datasets to define transcriptional states in adult human spinal cord central canal–associated cells. We show that progenitor-associated programs in intact tissue remain non-cycling and astrocyte-embedded, while proliferative and biosynthetic programs are unmasked under permissive ex vivo conditions — revealing a state-dependent rather than constitutive model of proliferative competence.
Rebuilding Spinal Circuit Computation Through a Patient-Specific Interneuron Precision Model
Preprints.org
We reframe spinal repair as the reconstitution of circuit computation, synthesizing how embryonic patterning programs establish interneuron diversity, connectivity, and network symmetry encoding motor coordination. Spinal cord injury disrupts this logic, fragmenting excitatory-inhibitory balance and desynchronizing rhythmic modules, while residual circuits retain latent capacity for resynchronization. We propose the Patient Specific Interneuron Precision Model (PIPM), a closed-loop framework linking patient-specific biological states — progenitor competence, morphogen sensitivity, and metabolic tone — to circuit-level computation and recovery potential.
Personalized Stem Cell-Based Regeneration in Spinal Cord Injury Care
International Journal of Molecular Sciences
A comprehensive review exploring the intersection of patient-specific variability, bioengineering innovations, and transcriptomic-guided precision medicine in spinal cord injury therapy. Covers transplantation of NSPCs, iPSCs, and MSCs, and how patient-specific factors — cellular senescence, genetic and epigenetic variability, injury microenvironment, and comorbidities — influence graft survival and differentiation. Discusses CRISPR-mediated hypoimmunogenic engineering and biomaterial-based delivery platforms for precision-driven SCI repair.
Transcriptomic and Functional Landscape of Adult Human Spinal Cord NSPCs Compared to iPSC-Derived Neural Progenitor Cells
Cells
The first direct transcriptomic and functional comparison of syngeneic adult human NSPC populations — bona fide spinal cord NSPCs versus iPSC-derived NSPCs regionalized to the spinal cord (iPSC-SC) and forebrain (iPSC-Br). RNA sequencing revealed that iPSC-Br NSPCs closely resemble bona fide spinal cord NSPCs, with enriched neurogenesis, axon guidance, and synaptic signaling pathways, while iPSC-SC NSPCs showed significant heterogeneity and suboptimal regional specification. Donor-specific factors significantly modulated transcriptomic profiles, highlighting the impact of genetic and epigenetic individuality on NSPC behavior.
These results were
produced by our platform.
The analytical workflows and computational approaches developed through these studies inform the design of the Boring Science platform. This is what boring looks like when it works.
Single-nucleus sequencing maps a non-proliferative progenitor state to astrocyte-rich regions of human spinal cord
Using snRNA-seq and spatial transcriptomics processed through our unified pipeline, we identified a distinct glial-enriched progenitor-like transcriptional program. UMAP clustering revealed progenitor-like cells (n=551, 0.0%) with significantly elevated astrocyte niche and ependymal scores. Spatial deconvolution confirmed localization of this program to astrocyte-rich tissue regions — linking molecular identity directly to spatial architecture.
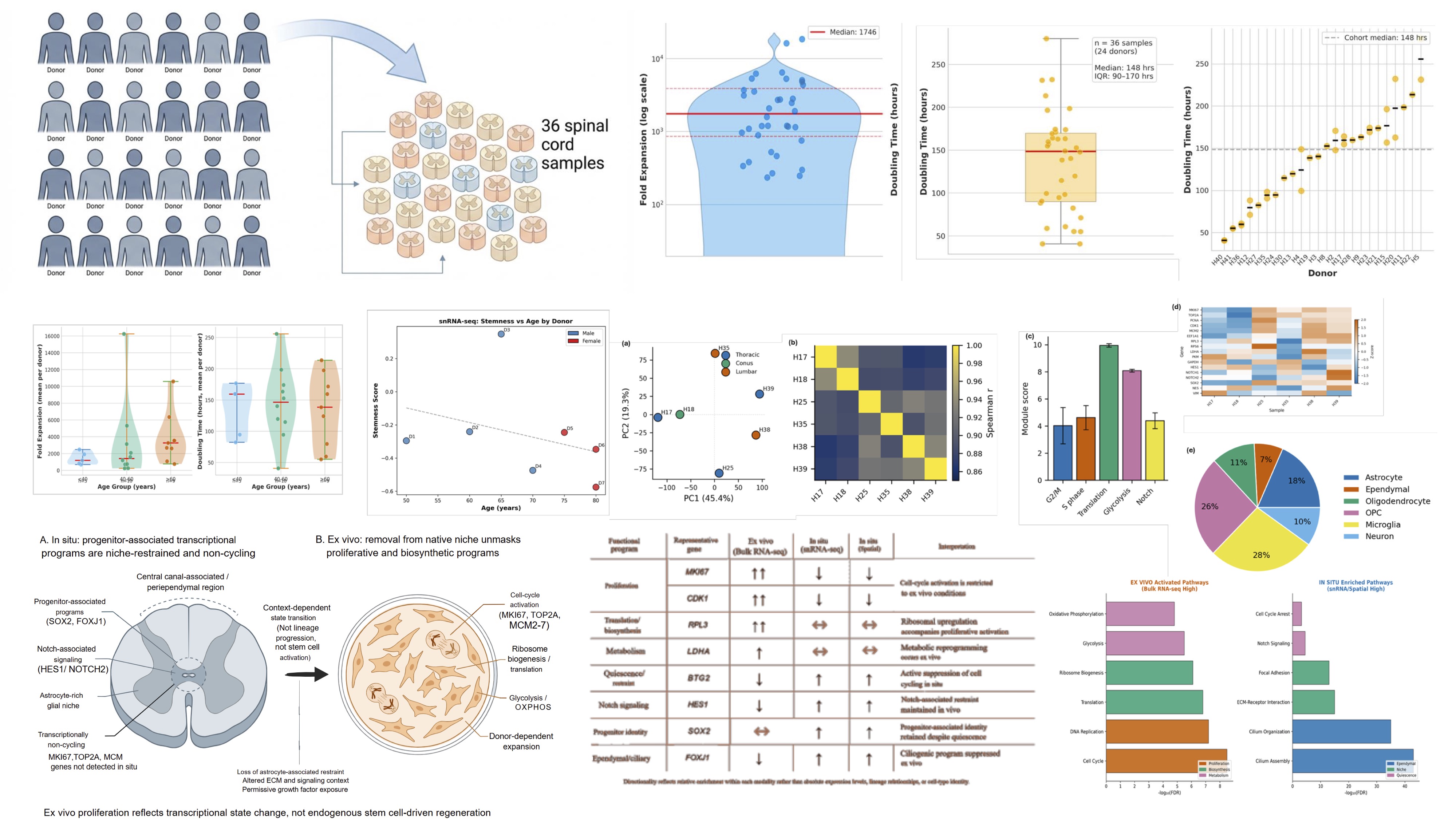
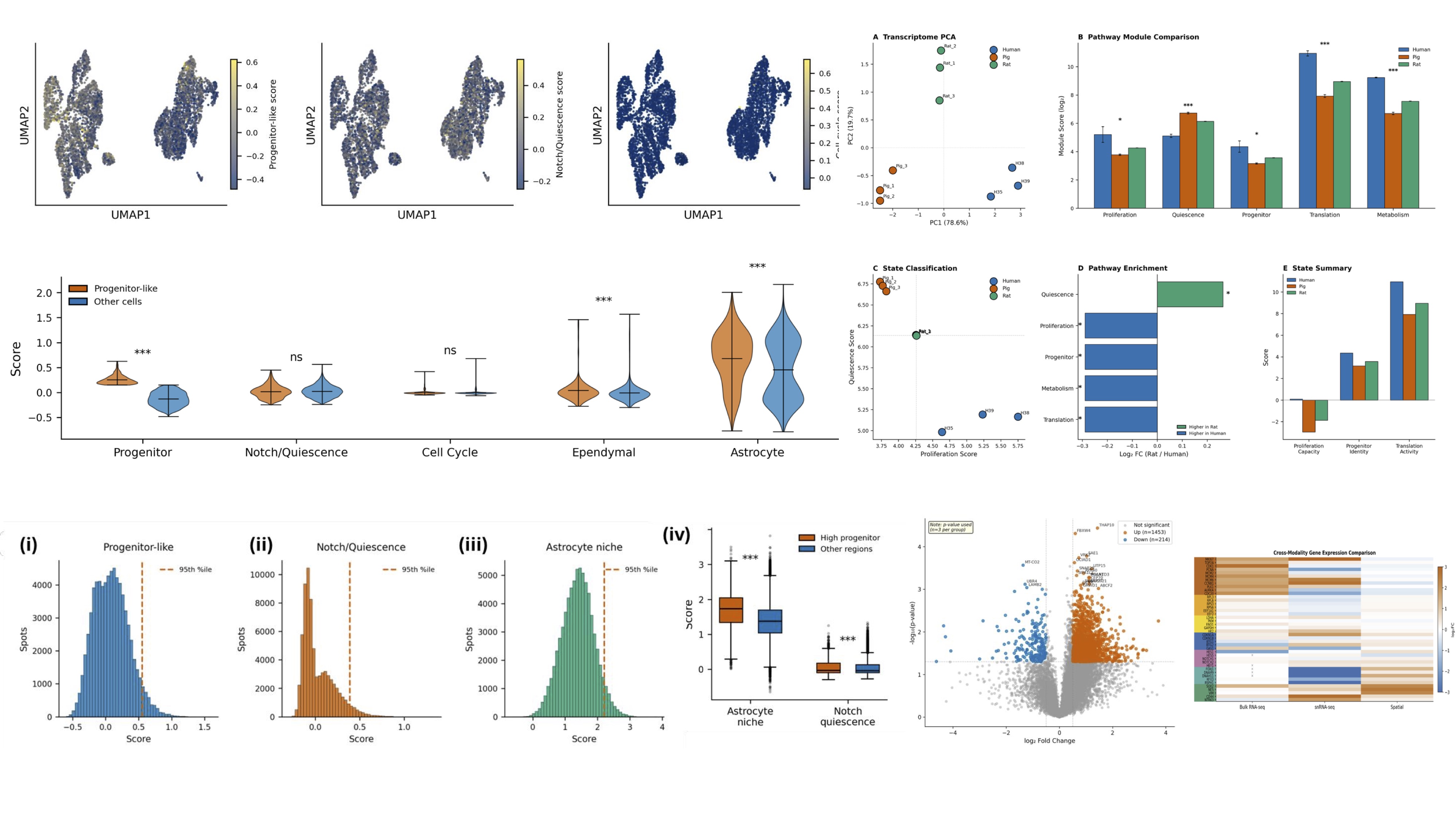
A unified cross-modal platform integrates snRNA-seq, spatial transcriptomics, and bulk RNA-seq to resolve cell state, tissue architecture, and cross-species identity in a single workflow
Our platform ran snRNA-seq, Visium spatial transcriptomics, and bulk RNA-seq data through a single reproducible workflow — enabling direct comparison across modalities without batch artefacts or manual harmonization. snRNA-seq UMAP resolved distinct progenitor-like, Notch/quiescent, and cell-cycle cell states; spatial scoring mapped these programs onto tissue architecture at spot resolution. Simultaneously, bulk RNA-seq processed across human, pig, and rat identified PC1 (78.6%) separating human from rodent, with pathway scoring confirming a high-quiescence, low-proliferation human phenotype. One platform. Three modalities. One coherent answer.
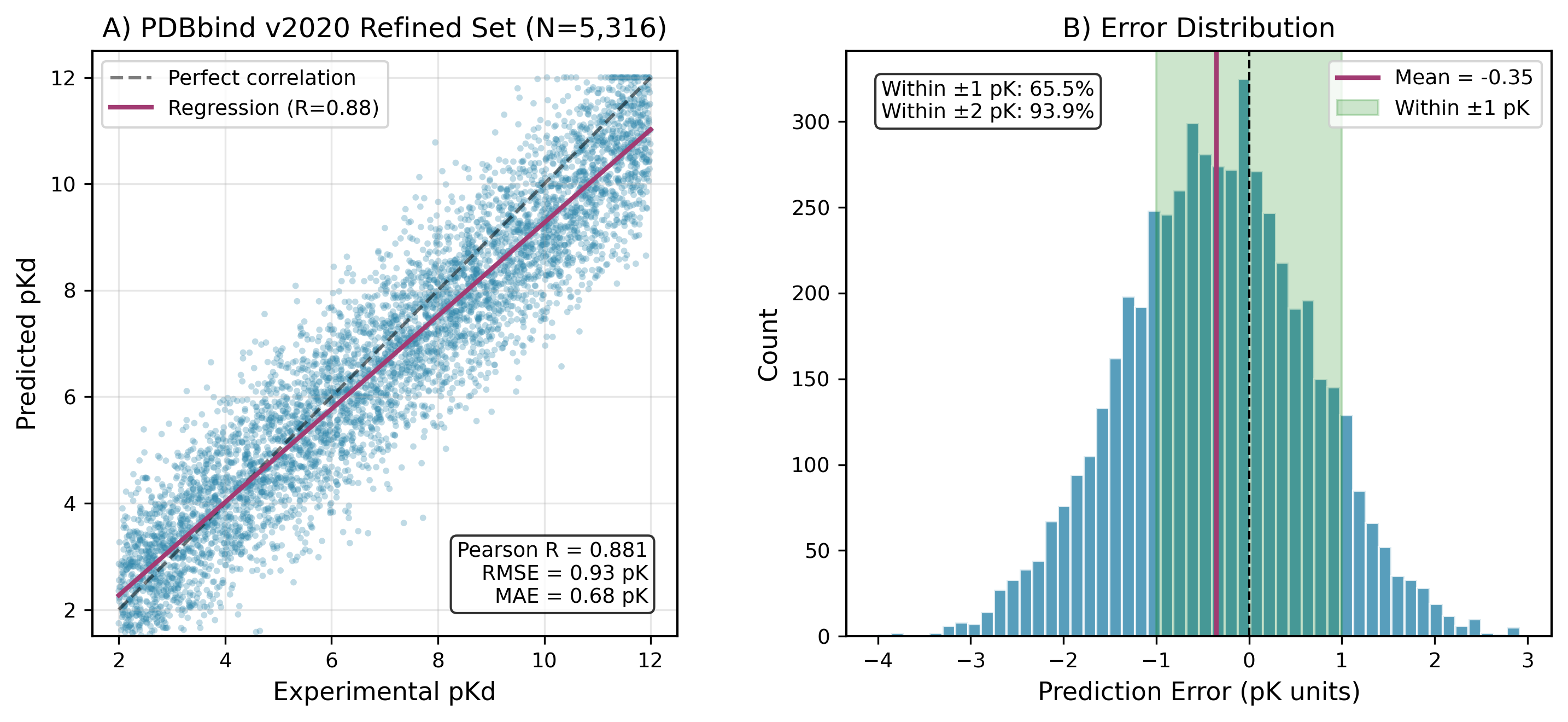
SE(3)-equivariant graph neural network scoring outperforms all classical docking methods — R = 0.88 on PDBbind, R = 0.82 on ULVSH, AUC-ROC = 0.94
bsdock encodes protein-ligand complexes as heterogeneous graphs with physically grounded rotational and translational invariance — a constraint that standard empirical scoring functions ignore entirely. Trained on 5,316 PDBbind complexes and 942 ULVSH compounds, the SE(3)-GNN achieves 5.5× higher affinity correlation than the best classical baseline (VM2). AutoDock Vina, Gnina, MM-GBSA, and MMPBSA all cluster near zero correlation on the ULVSH benchmark. The hybrid workflow — traditional pose generation followed by GNN rescoring — completes a full docking run in under one minute. Generalisation to a novel GABA receptor series (R = 0.68) without target-specific training confirms the model learns transferable binding physics, not dataset-specific patterns.
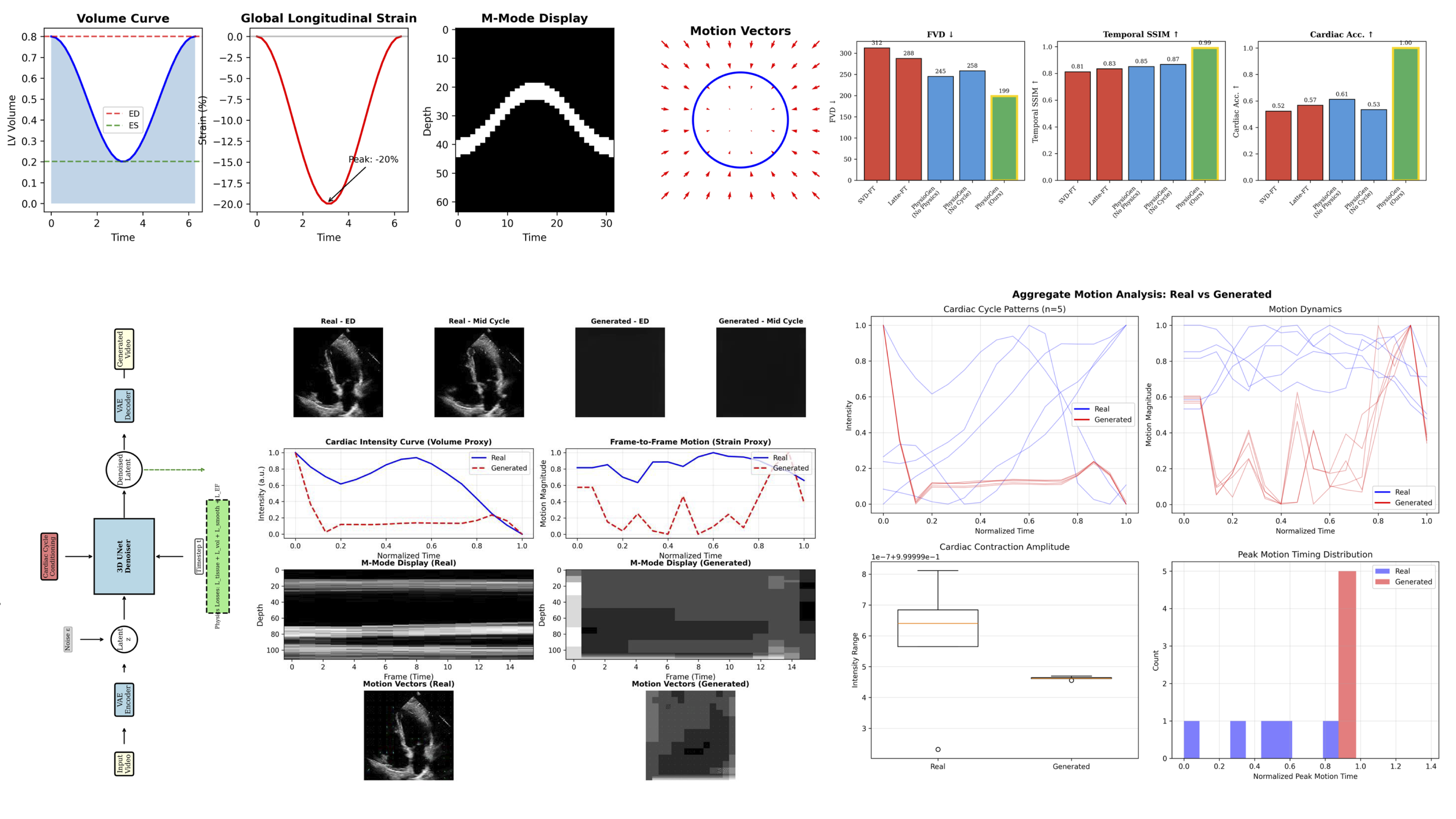
Physics-constrained video diffusion achieves SSIM = 0.87 — a 6.2× improvement over baseline — with 12.8% downstream gain in ejection fraction prediction
Standard video diffusion models optimize perceptual quality without enforcing the physics of biological motion — producing videos that look plausible but violate cardiac biomechanics. bsvision addresses this with a pixel-space 3D UNet conditioned on cardiac cycle embeddings (sin/cos phase encoding), where cross-frame attention at all encoder/decoder levels allows every frame to attend directly to all 16 frames in the sequence. Five ablation iterations isolated the dominant contributors: cross-frame attention adds +0.32 SSIM, SSIM loss weight tuning adds another +0.32, while physics-based loss terms (temporal smoothness, flow consistency, jerk penalty) contribute +0.04 SSIM as regularization. The final model achieves SSIM = 0.87 between consecutive frames — 6.2× over V1 baseline (0.14). Augmenting downstream models with 10k synthetic samples from bsvision yields a 12.8% improvement in ejection fraction prediction accuracy, validating the clinical utility of the synthetic data.
Doing interesting
work? Let's make it boring.
We're open to research collaborations, co-authorship, and data partnerships.
inquiries@boringscience.bio